|
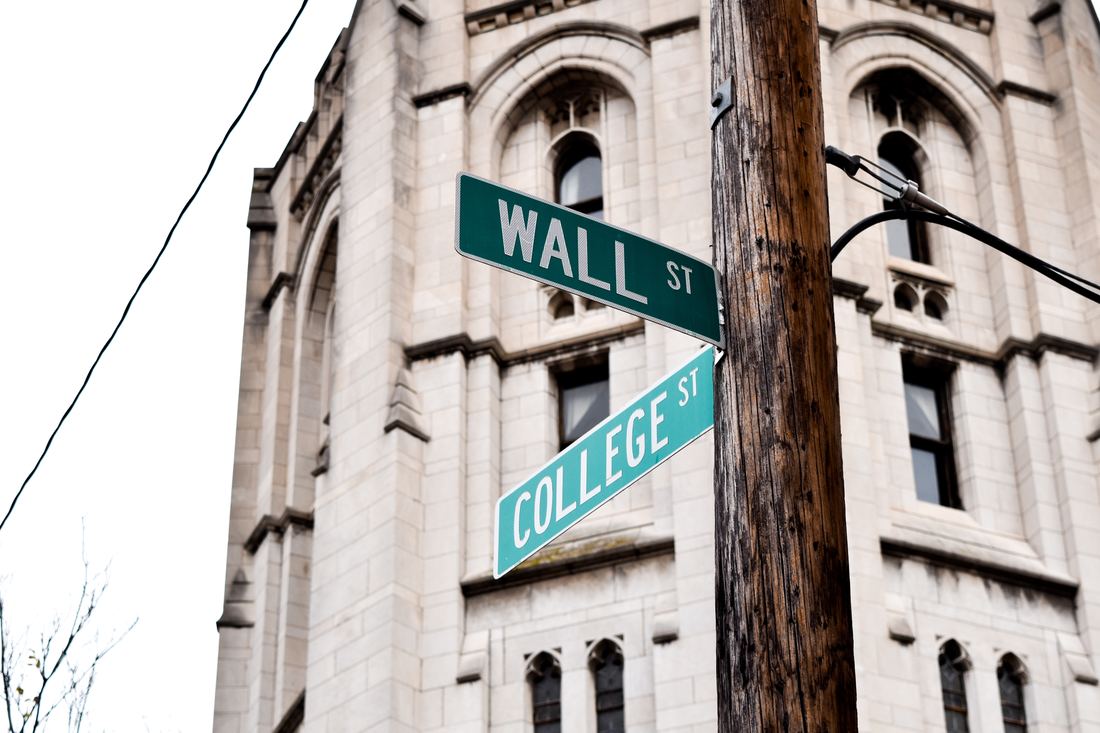
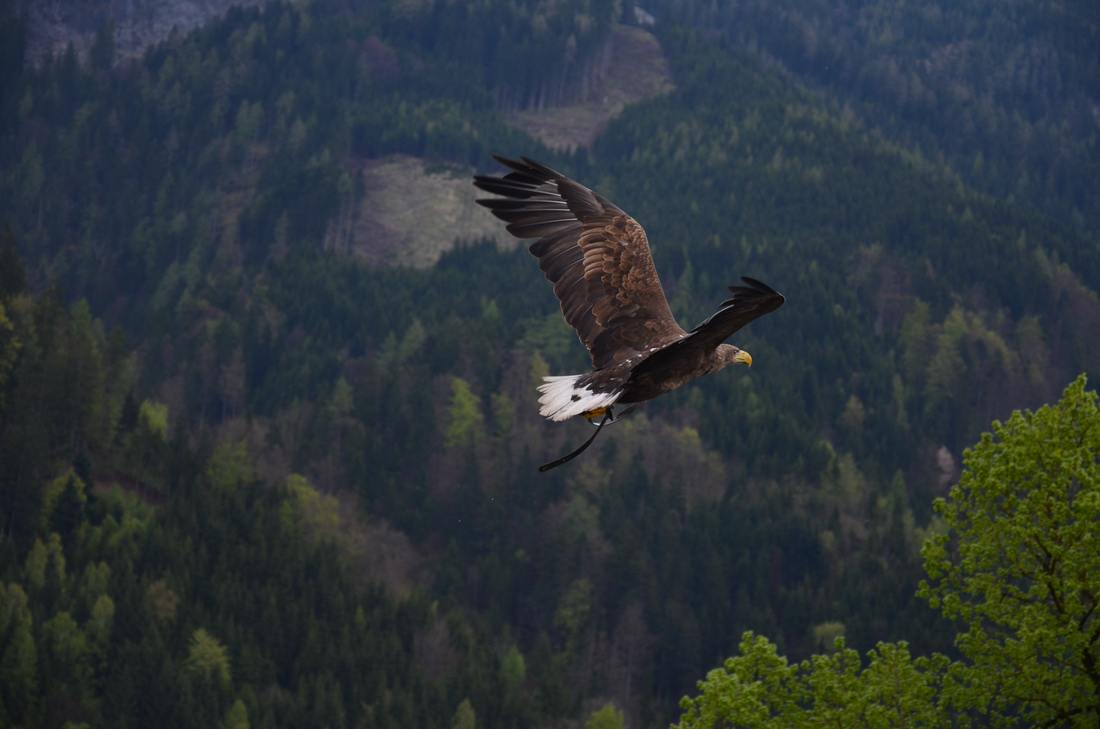
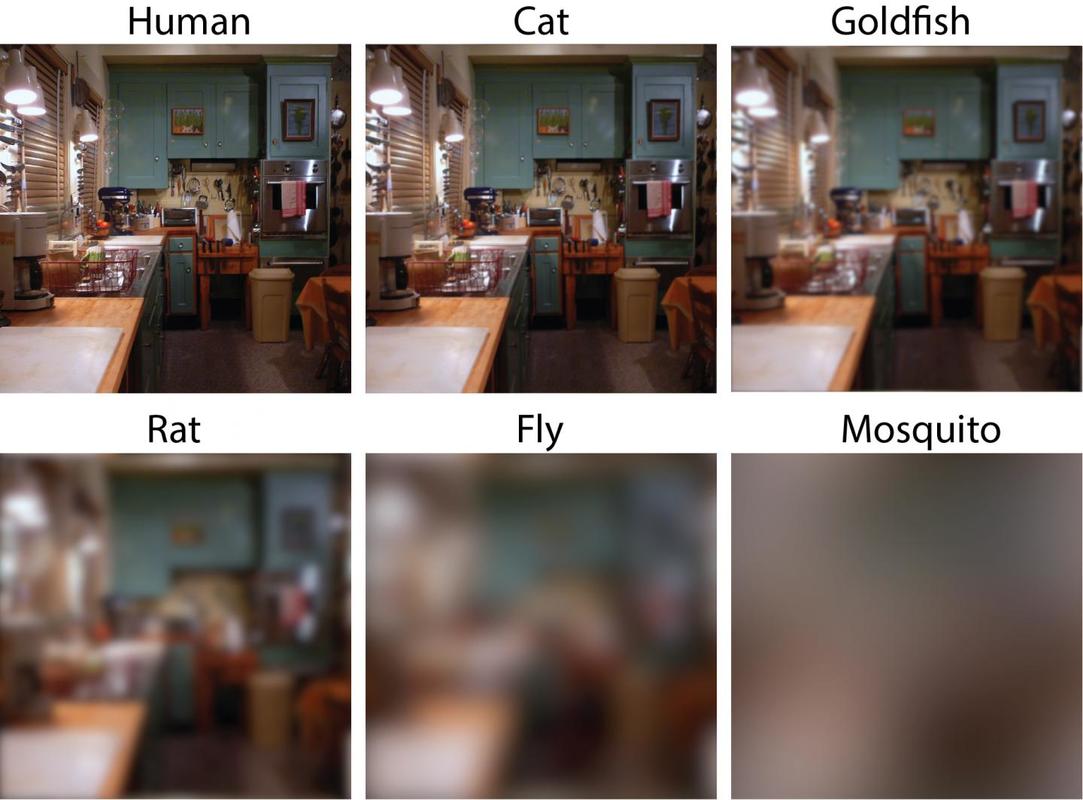
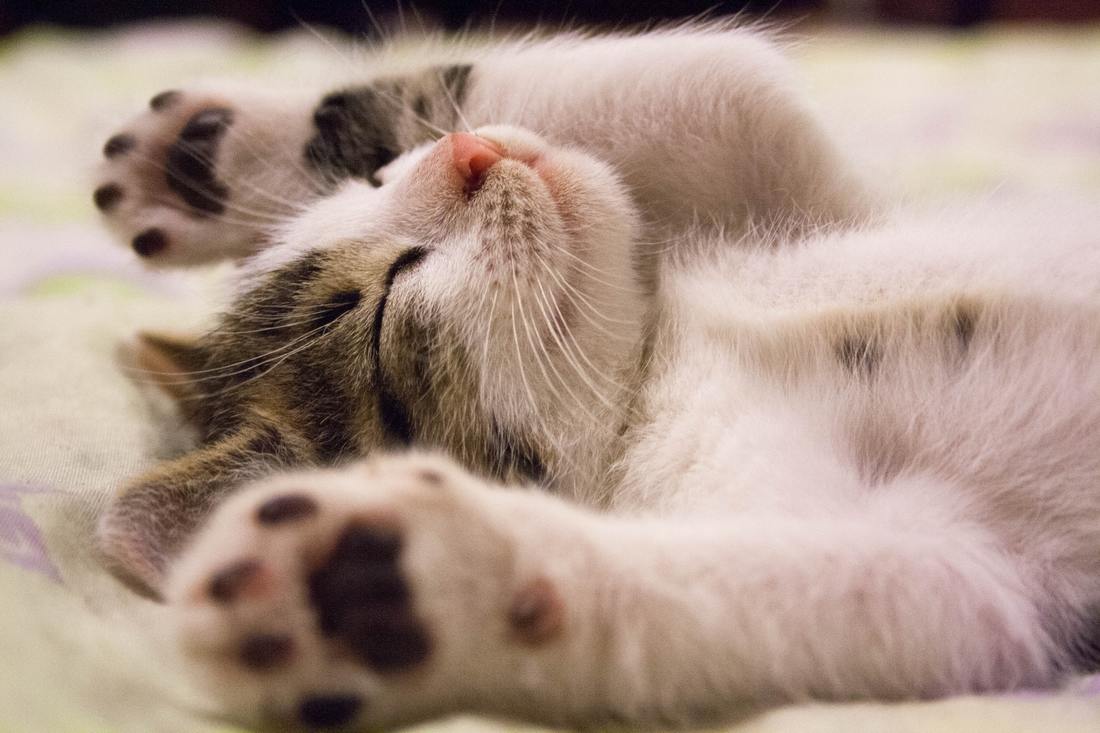
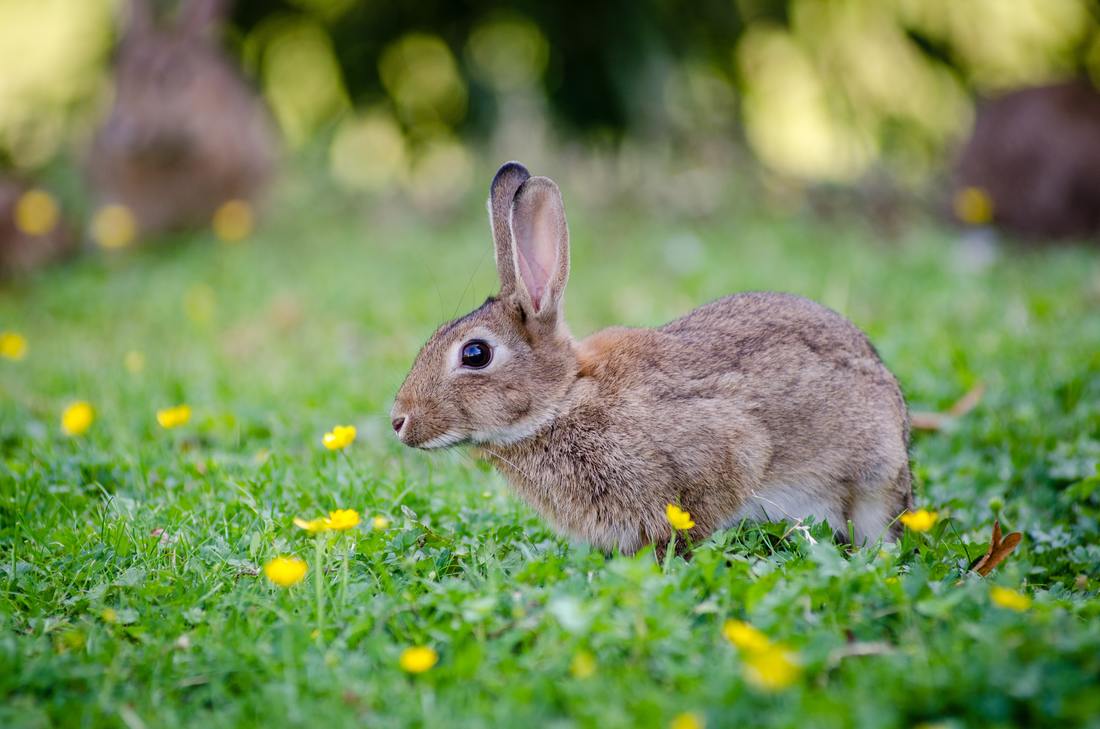
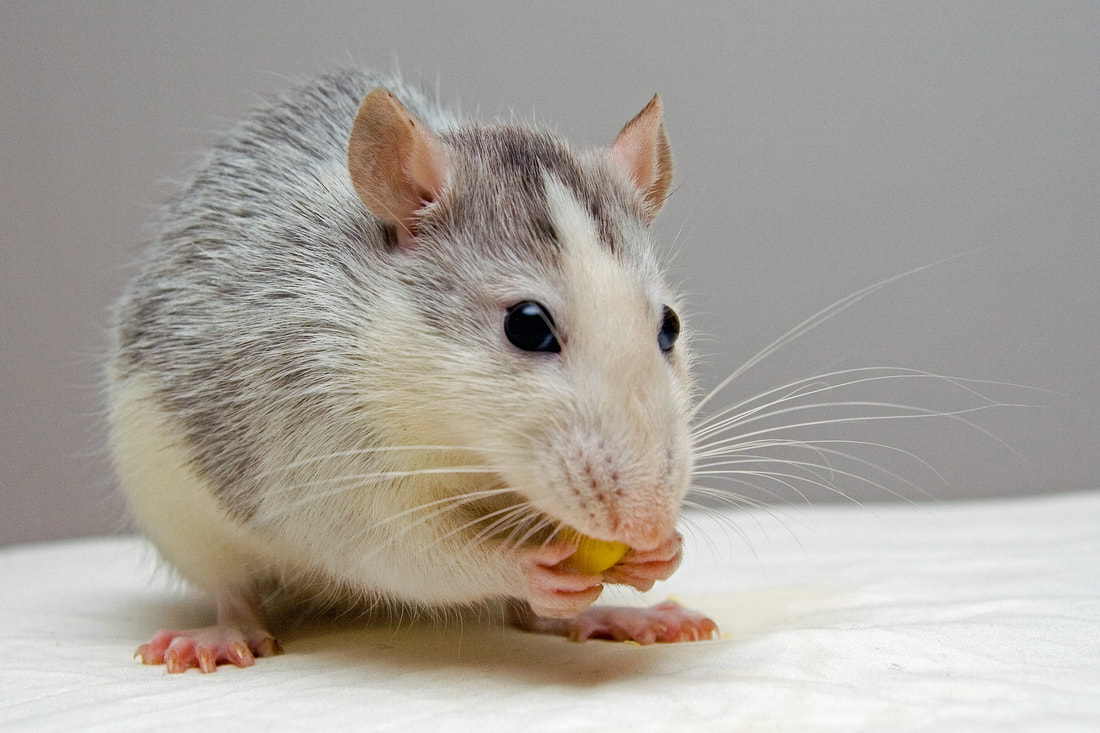
While you might have heard that animals don’t necessarily see the same colours as us (spoiler alert, dogs don’t actually see in black and white), have you ever wondered if they can see things as clearly as we do? How clear or blurry something appears has to do with visual acuity. Visual acuity refers to how finely or precisely our eyes are able to perceive information, and this determines how well we can process visual information. Imagine trying to read a street sign from a long distance. It’s our visual acuity that determines whether we’re able to read it from fifty metres away, or twenty, or ten. While we might assume that animals have the same visual acuity as us, this is actually not the case. Scientists have estimated that visual acuity spans over four orders of magnitude in the animal kingdom. That means the most sharp-sighted animal has a visual acuity that is ten-thousand times better than the one with the blurriest vision. A recent review, published in Trends in Evolution & Ecology examined the visual acuity of a range of species, collated from a number of different scientific studies. Visual acuity was measured in terms of cycles per degree, which is how many pairs of black and white parallel lines a species can discern within one degree of the field of vision before they turn into a big smudge. Since it’s a bit hard to get a fish to sit down in front of an eye chart and tell you what it sees, the researchers typically used behavioural tests or special cameras and imaging to examine the animal’s eyesight. It is possible to estimate visual acuity based on a number of factors within the eye, such as the focal length and receptor diameter. Humans are pretty high up on the visual acuity ladder and are able to discern approximately 60 cycles per degree. Other primates, like chimpanzees scored about the same. Only some birds of prey scored higher, like eagles that can spot a tasty rabbit snack from hundreds of metres away. For the most part, birds scored pretty low, with a median acuity of 11 cycles per degree. In human terms, vision of less than 10 cycles per degree is considered to be legally blind. Interestingly, visual acuity can vary a lot between predators and prey, and this may explain some of the unusual patterns exhibited in the animal kingdom. The researchers developed software that could simulate the visual acuity of various species (important note – this does not necessarily reflect what the animal is actually ‘seeing’ because the information sensed by the eyes still needs to be processed). They looked at images of Orb-Weaver spider webs, and discovered that they were visible to birds, so they could avoid flying into them, but invisible to the spider’s insect prey. They also looked at butterflies spots – which have sometimes been thought to send messages to other butterflies – and realised that it was unlikely the butterflies could even see them. But apart from explaining why spiders have cool webs or why some butterflies have spots, understanding visual acuity in animals can have huge implications for animal handlers and trainers. When training an animal we often assume they’re able to see what we can, but this review shows that’s not necessarily the case. Unfortunately, many domesticated animals like dogs, cats and horses were not covered in this study, but the majority of species examined fell in a range of what would be considered legally blind to humans. And while visual acuity isn’t the be-all and end-all to animal vision, since a lot of post-processing occurs in the brain, the point is that most animals don’t see the same way as we do. So next time you’re interacting with an animal, take a moment and try to consider things from their point of view.
0 Comments
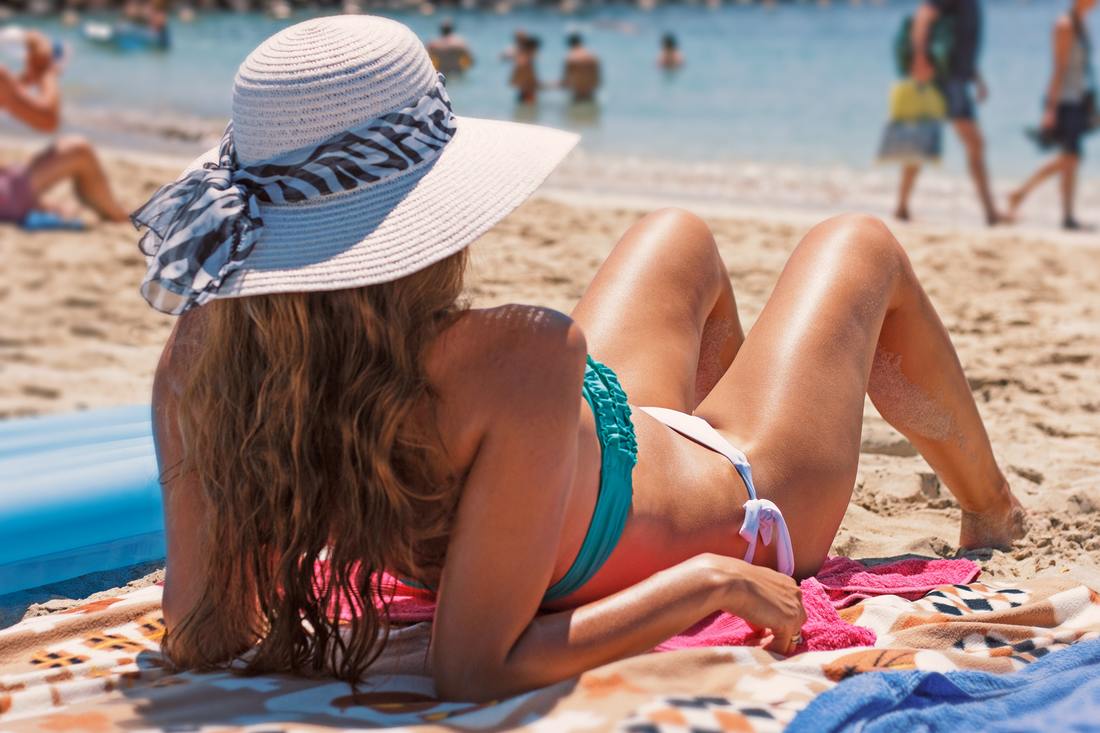
Ever wondered why some of your friends seem to get a deep tan over summer while you get burnt to a crisp after only minutes in the sun? It could be due to your genes. Previous studies have shown that the heritability of ease of skin tanning is about 45%. That means if both your parents tan easily, there’s a good chance you will too.
While some of the genes associated with skin pigmentation and tanning have been identified, a recent study published in Nature Communications has identified 10 new genes connected to ease of tanning, as well as confirming 10 previously known genes. In order to find these new genes, the researchers analysed a genetic database of over 120,000 individuals who were divided into two groups based on their ability to tan. This analysis revealed 30 different genes that may be involved, and these were cross-referenced with other large databases to determine 20 genes linked to tanning, 10 of which were novel associations. But why do tanning genes matter? Well, as we all know, sun exposure can cause skin cancers. Additionally, people who burn more easily are at greater risk. While some of the genes have previously been associated with skin cancer, this study has identified new genes that could affect cancer risk. There wasn’t enough data in this study to determine how well ease of tanning correlates with melanoma risk, but the researchers did find a genetic correlation between ease of tanning and non-melanoma skin cancer. The same genes that were involved in decreased tanning ability increased the risk of non-melanoma skin cancer. These findings are important to help identify people who may be at greater risk of skin cancer. But don’t be complacent if you are someone who tans easily – previous work has shown that people with darker skin who also carry lighter-skin genes have a similar risk of developing skin cancer as people with light skin. The researchers hope to continue their work to further investigate how these genes contribute to tanning and skin cancer, and to assess the degree to which they affect a person’s risk of developing skin cancer. However, no matter what’s in your genes, remember that no tan is a safe tan and the best way to minimise your risk of cancer is to slip, slop, slap and avoid sun exposure. How many times have you woken up thinking ‘what a strange dream’ but have then been unable to recall it later during the day? And how many times do you wake up thinking you haven’t dreamt at all? Dreams can occur during any stage of sleep, but mainly happen during rapid-eye movement (REM) sleep when the brain is most active. It is estimated the average person dreams 4-6 times per night. No-one knows for sure why we dream, but it is thought of as a way for our brains to process the events of the day. Lucid dreaming, sometimes called a ‘waking dream’, is a type of dream where the subject realises they are dreaming and is able to control the dream. Apart from being a lot of fun, lucid dreaming is an active area of research for psychologists, as it could be used as a way to control or prevent nightmares, or provide therapy for PTSD patients.
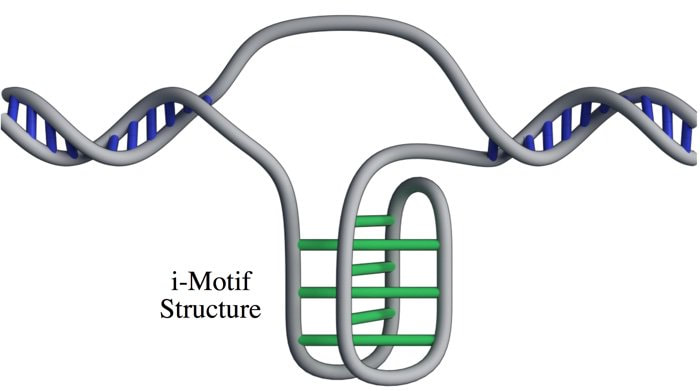
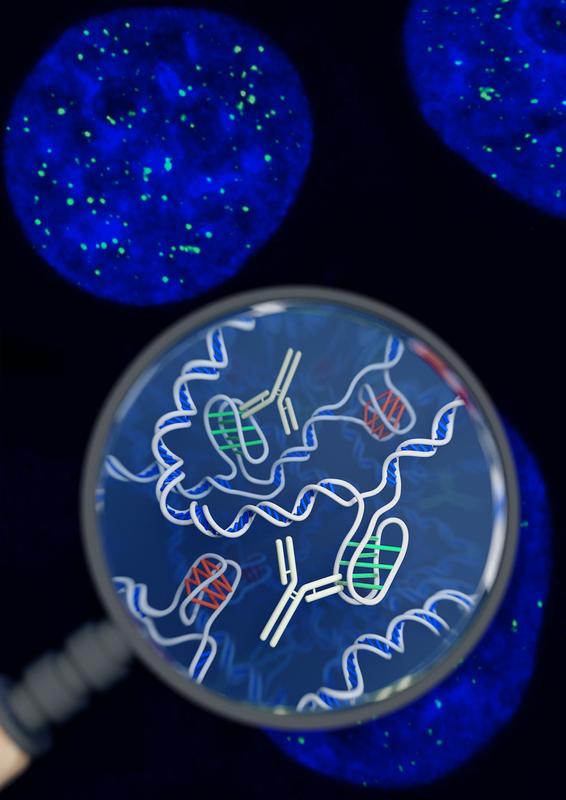
Lucid dreaming is actually a skill that can be taught or developed. But the first step to lucid dreaming is being able to remember your dreams. A recent study by researchers at The University of Adelaide has found that taking a vitamin B6 supplement before bed could help you to remember your dreams. The randomised, double-blind, placebo-controlled investigation (that means it’s pretty good in terms of scientific studies) involved 100 participants. The study found that participants who took a 240 mg supplement of vitamin B6 before bed experienced 61% better dream recall than the control group over the 5-day period. It’s still not clear why this effect occurs. One theory is that it happens because vitamin B6 plays a role in converting the amino acid tryptophan into the neurotransmitter serotonin, which increases time spent in REM sleep. An alternative theory is that vitamin B6 disrupts sleep, so sleepers are more likely to wake mid-dream and remember their dream. However, no significant effect on time awake during the night or sleep quality was observed in this study. Participants during the study were advised to avoid foods naturally high in vitamin B6, like wholegrain cereals, milk, cheese, legumes and eggs, so it is unclear as to whether this boost in dream recall only occurs if there is a deficiency in vitamin B6. And, if you want to try this for yourself, the researchers warn that taking vitamin B6 supplements before bed may exacerbate existing REM sleep conditions, and the effects on dream recall may diminish over time. In the future, the researchers hope to investigate the validity of the serotonin theory, and determine if the effects of vitamin B6 vary according to how much is included in the diet. If you want to try this for yourself, or learn more about lucid dreaming, you can check out the researcher’s website at www.luciddreamingaustralia.com. Most of us are pretty familiar with the double-helix structure of DNA; the one that looks like a spiral staircase with two strands of DNA pairing to each other. But did you know DNA can come in lots of other shapes too? DNA can twist and turn in different directions, pair up differently, and even form a triplex structure. Studying the structure of DNA is important because it gives insights into how DNA is used and read within the cell. Our DNA is really long, so it has to be packed in pretty tightly to fit inside the nuclei of our cells. In order to read the DNA to express genes it has to be unpacked and unwound, and there are lots of different proteins and big molecules (like RNA) that come into play in this process. Understanding the different structures of DNA and how they regulate gene expression can tell us information about what is happening in the cell and how that might impact health and disease. One structure – the i-motif – looks a bit like a knot of DNA. Normally the DNA helix structure is formed by the pairing of base letters A, T, C and G on two different strands of DNA, but the i-motif ‘knot’ structure is formed by the C-bases on the same strand of DNA binding to each other. (Zeraati et al., Nat Chem, 2018) This had only ever been observed in vitro, that is, outside of the human body, and usually only under harsh pH conditions. While there was some indirect evidence for its formation in vivo, it wasn’t known whether or not it actually existed in living organisms. This week, Australian scientists announced they had found a way to detect this i-motif inside living human cells for the very first time. Their findings were published in the journal Nature Chemistry. The researchers developed an antibody that binds specifically to the i-motif structure. They did this through three rounds of identifying and refining a mutant antibody that could bind to a known i-motif structure found at the end of chromosomes. They found the antibody bound strongly and specifically – that is, it didn’t interact with other proteins, RNA, or other DNA structures. They tested the antibody on six other known i-motif structures and found it was able to bind to all of them, indicating the antibody can bind to i-motifs regardless of the DNA sequence. That meant it could be used for detecting i-motifs inside cells. The researchers stained human cells using a technique that makes fluorescent spots in places where the antibody binds. They found multiple fluorescent green dots, showing for the first time the presence of i-motifs in living cells. They observed the i-motifs appearing and disappearing over time as the cells grew and divided, and found they were linked to regulatory regions of DNA. This research not only gives direct evidence for i-motifs in cells but also provides preliminary evidence that they could be involved in human diseases that arise from dysregulation of gene expression, such as cancer. Mutations in i-motif regions could change the stability or shape of the i-motif and affect how it interacts with DNA. This research, and the new antibody tool, opens new doors for future studies to explore the biological roles of i-motifs and potentially develop new therapeutics to treat disease. Artist's impression of the i-motif DNA structure inside cells, along with the antibody-based tool used to detect it. (Chris Hammang).
With a quarter of the year gone it's time to reflect back on our New Year's Resolutions. Mine was to blog every week, and so far I've done alright. The last month or so has been pretty busy which hasn't left much time or energy for writing but I think I've only missed a week or two (not including that one time I got lazy and reposted something I wrote last year).
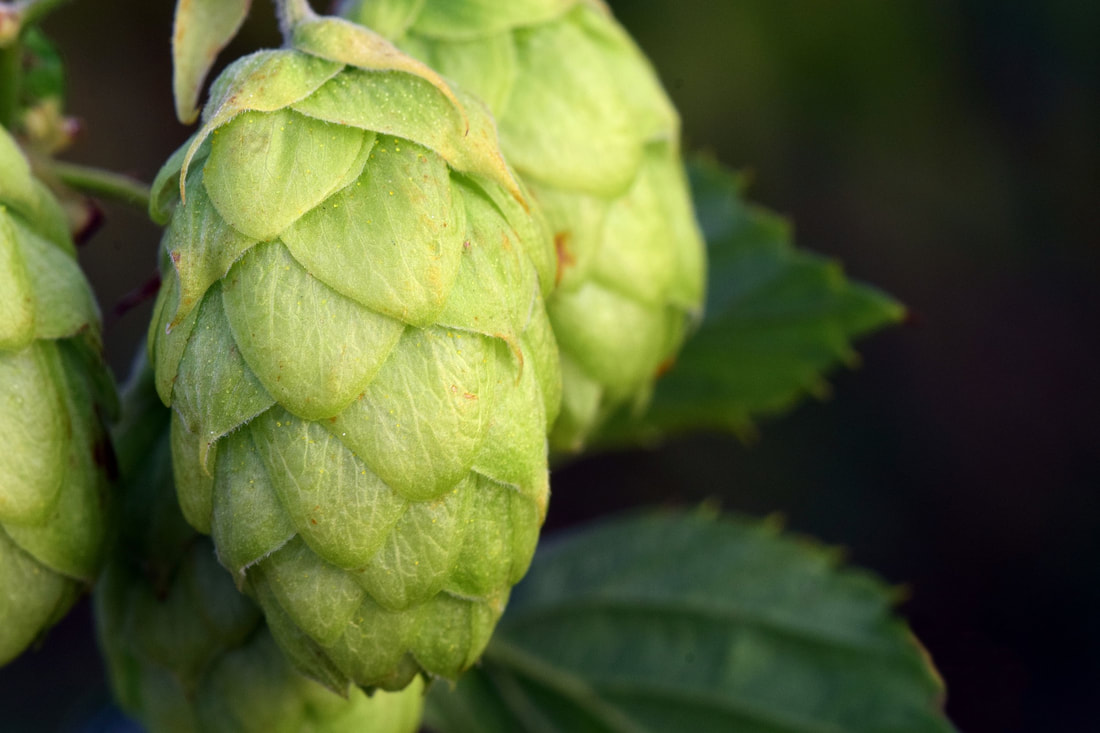
On that note, I'm just checking in to say Sci Chat will be taking a break for a few weeks while I move to Canada to start my Endeavour Fellowship. I'll be conducting research towards my Masters degree over at the University of Calgary for 6 months. I'm leaving this Thursday (April 5) and will be travelling a little before starting my research on April 16. Give me a few weeks to get settled in and then Sci Chat will be back up and running by late April. Until then, thank you to my regular readers. I hope you're enjoying my blog and I'll be back soon! Craft beers have become wildly successful over the last few years and one reason consumers are so crazy about them is that key ingredient – hops. Hops are a plant added during the beer making process to impart a bitter flavour to balance out sweetness, and add unique tropical or citrus aromas or flavours. But all those darn hipsters and their hoppy beer means the global supply of hops can’t meet the demand, making them very expensive. On top of that, they require a lot of water and natural resources to grow. Furthermore, the actual content of the essential oils within the hops can vary wildly, so it’s hard to achieve a consistent hoppy taste in beer. Researchers at the University of California – Berkeley have genetically engineered a strain of brewer’s yeast that means you can make hoppy beer without the hops.
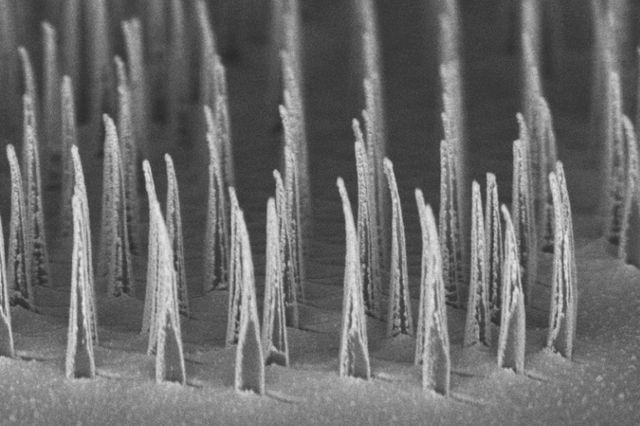
First they identified two particular chemical compounds that give hoppy beer its special taste and smell, and then they found enzymes (biological catalysts) that can make these compounds from precursors already produced naturally by the yeast. They used CRISPR/Cas9 gene editing technology (you know, those molecular scissors I talked about in a previous post?) to genetically engineer yeast to produce high levels of these flavour molecules. This is a pretty impressive achievement because firstly, brewer’s yeast, unlike domesticated wine yeast, is pretty wild and has lots of crazy things going on with its chromosomes, making it notoriously difficult to genetically manipulate. Secondly, making sure those genetic manipulations stick so that the population is stable is pretty challenging. The researchers then used these new strains of yeast to ferment beers and checked that the levels of the flavour molecules were consistent. They showed much less variation than traditional dry-hopped beers, which will help brewers ensure a consistent taste. But, as they always say, the proof of the beer is in the drinking. A panel of taste testers consistently rated the beers as having a hoppy flavour and aroma. Of course, the complex flavour of hops cannot be completely encapsulated by just two molecules, but these methodologies provide a foundation to build more complex yeast strains that produce a wider variety of flavours and aromas. Ultimately, this technology means not only can you brew hoppy beer without the hops, but it paves the way for further improvements in brewer’s yeast and advancements in environmental sustainability. Many human diseases such as cystic fibrosis, muscular dystrophy, AIDS, and cancer are caused by disruptions to genes. Gene therapy is an emerging technology that aims to replace missing or defective genes within human cells to treat these conditions. However, it is currently still very risky and not widely tested on humans. Existing approaches to gene therapy use viruses, electric fields or harsh chemicals. These may pose a risk to the cell in terms of toxicity or undesirable off-target effects, as well as being expensive and potentially inefficient. Scientists are currently working on ways to make gene therapy safer, faster, cheaper and more efficient. One of the major challenges in delivering gene therapy is making sure the therapeutic agents can target the desired areas and enter the cells. The trick is to get the therapeutics to enter the cell without killing it or affecting its growth. A group of researchers at the University of California – Los Angeles have developed tiny ‘nanospears’ made of silicon, nickel and gold. Because the tips of these spears are so infinitesimally small they are able to penetrate the membrane of the cell without harming it. Nanospears aren’t new – other groups have worked on similar structures powered by nanoscale motors in order to propel the spears towards their target. However, these newly-developed spears are even better as they are magnetic, so scientists can use a magnetic field to guide the nanospears without having to use toxic chemicals. They are also bio-degradable and can be mass-produced, making them cheaply available on a large-scale.
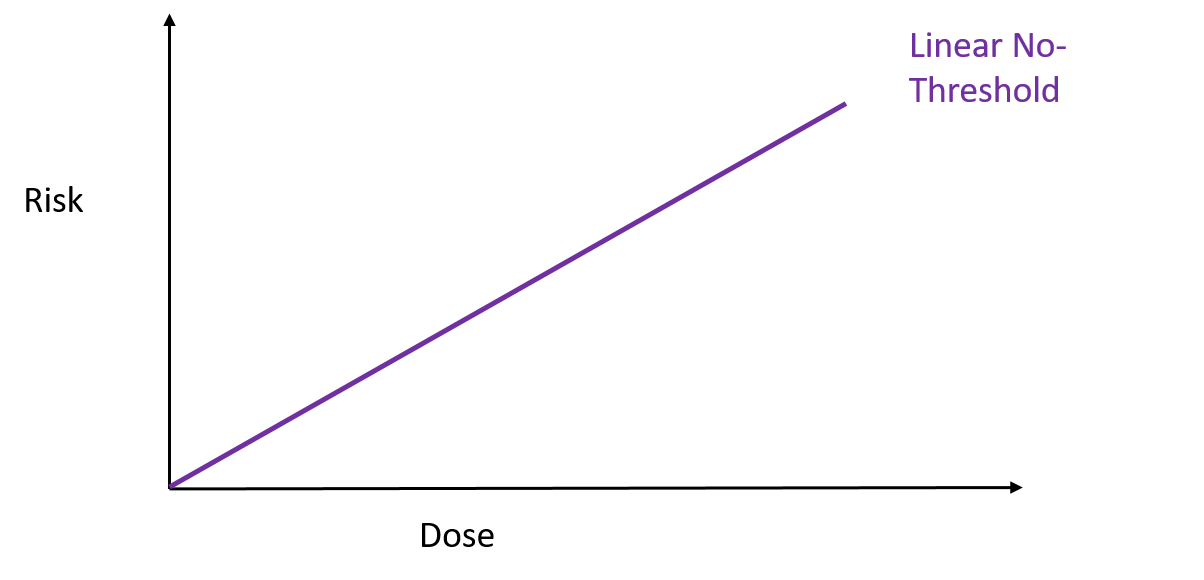
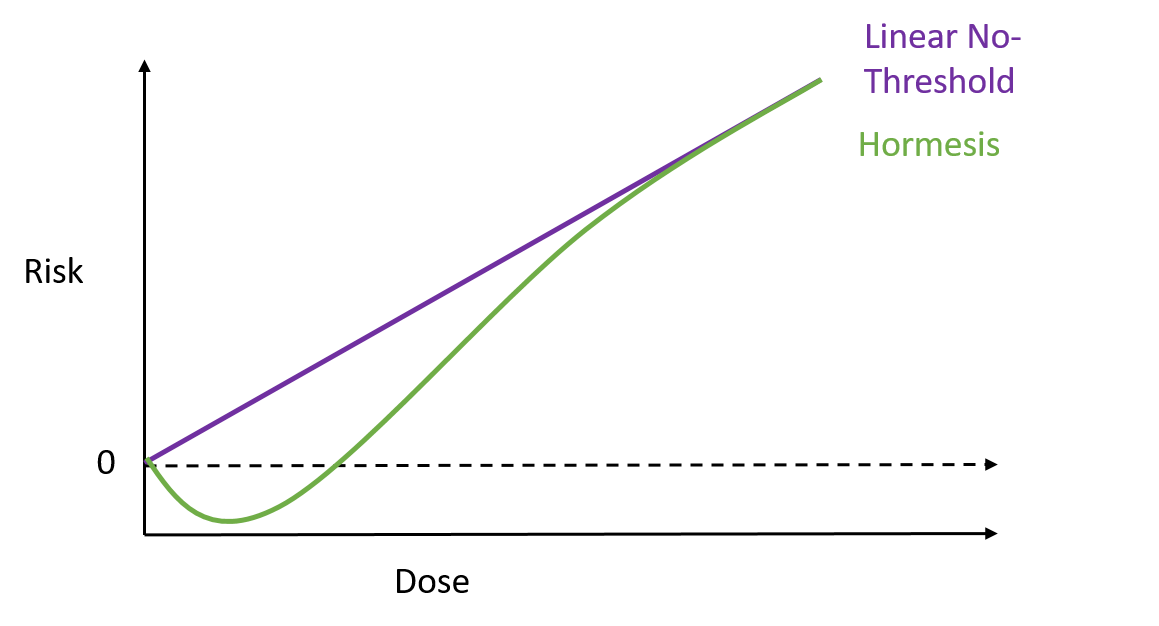
To test how well these nanospears worked, the researchers used them to deliver a gene for green fluorescent protein – that’s right, it does exactly what it says on the label – into a culture of human cells. Approximately 80% of the cells produced a green glow, indicating the gene reached its target, and of those cells 90% survived. That’s a marked improvement over current forms of gene therapy. So far this technique has been used to deliver DNA to cell culture systems. Not only is this an extremely a useful tool for researchers, but it may also help speed up the delivery of gene therapy and immunotherapy. But that’s not all. These nanospears could be used to deliver more than just DNA, paving the way for new nanomedicine technologies that could be used for applications ranging from biological research to human medicine. The researchers are currently working on optimising the nanospears technology and hope that in the future it could be used to guide the delivery of gene therapies within a human patient. Hey everyone, this week's blog is a bit different to normal because it's based on a body of research rather than a single recent paper. I'll admit, I got a bit behind this week so I've recycled a post from my Sci Comm class last year. It's a little longer than normal but well worth the read in my opinion - let me know what you think! Radiation can be a pretty scary thing. After all, the effects of an overdose of radiation can be devastating. Radiation overexposure can cause radiation sickness, burns, cataracts, infertility and cancer. Some of these effects can take years to manifest. The reason radiation is so dangerous is because it is able to damage cells by disrupting the chemical bonds between molecules inside the body. If a cell is greatly damaged it will die, and the death of many cells can lead to radiation sickness and even death. Alternatively, if radiation damages a cell’s DNA it can cause it to mutate and become cancerous. But radiation is an everyday part of life. On a daily basis we are exposed to radiation from space, air, rocks, buildings, food, and even from our own body. Radioactive elements are everywhere, and there’s no way of avoiding them completely. What matters is the dose Generally we think of radiation exposure as directly dose dependent -- that is, any radiation is harmful, with no safe level of exposure. This is called the linear no-threshold model and this is the model used by regulators and policy-makers to determine radiation guidelines. However, there’s evidence that this might be wrong. While there’s no question that high levels of radiation leads to a greater cancer risk, there’s less data about how low levels of exposure might affect this risk. This is partly because our natural incidence of cancer is fairly high and can vary greatly depending on lifestyle factors, making it hard to untangle the exact effects of chronic low-dose radiation. Despite this, there are some studies that show exactly the opposite of what you would expect: that low doses of radiation might be good for you. In Taiwan, during the 1980s, a batch of recycled steel was accidentally contaminated with radioactive cobalt. This steel was used to make over 180 buildings containing roughly 1700 apartments. Approximately 10,000 residents of the apartments were exposed to higher than normal levels of radiation over 9-20 years. Within this group of residents, the cancer mortality rate was much lower than in the general population – in fact, it was only 3% of the expected rate. The rate of congenital birth defects was also significantly lower – only 6.5% of the expected rate for the general population. This is possibly the best example of a case study on low-dose radiation because the sample size is large, and other factors such as lifestyle and environment are likely to be consistent between the sample population (the residents) and the general population of Taiwan. But it is not the only example. A number of other studies have shown that people living in areas with naturally higher background radiation have a lower cancer mortality rate than the general population. So, it appears chronic low doses of radiation may actually lower the risk of cancer. This phenomenon is termed ‘radiation hormesis’. It’s not certain why radiation hormesis may exist, but several explanations have been proposed.
It’s possible that radiation hormesis is an adaptive response to damage. Cell exposed to a very low dose of radiation, given a few hours to recover, and then exposed to a higher dose of radiation are less likely to sustain DNA damage than unexposed cells. This means cells that are pre-exposed are more able to protect themselves against higher levels of radiation. Other effects of low dose radiation include:
Animal studies have shown support for radiation hormesis. One experiment using mice predisposed to cancer found low doses of radiation helped slow down the rate of cancer growth. However, analysis of data in humans suggests a mixture of effects. Phillipe Duport, the founder of the Centre for Low Dose Research at the University of Ottowa says “A little radiation may be good for some people but bad for others.” Some scientists believe there is already enough evidence to accept the radiation hormesis hypothesis, and go so far as to say the linear no-threshold model is ‘scientifically unfounded’. Others believe proponents of the hormesis theory have chosen to ignore works that support the linear no-threshold model. However, most of the data supporting the linear no-threshold model is from survivors of the Japanese atomic bomb. These results may be complicated by other factors, such as exposure to cancer-causing chemicals released during the bombing. The conflicting evidence demonstrates a need for further research on humans, but this poses an obvious ethical dilemma. A strong experimental design would require a large number of subjects willing to be exposed to radiation over a long period of time. Unfortunately such a study is unlikely to be approved by an ethics committee, so we may have to rely on data from ‘happy accidents’ like the Taiwan apartments. Currently, advisory bodies like the National Council on Radiation Protection and Measurements are sticking to the linear no-threshold model. On one hand, acceptance of the hormesis model would redefine ‘safe’ levels of radiation exposure, and potentially save millions of dollars in the clean-up of radiation accidents. On the other, regulators tend to err on the side of caution, and mistakes in this area could be devastating to the lives of the people affected. Nevertheless, there’s stacks of evidence to support radiation hormesis, and pressure is growing on policy makers. So perhaps we will see a change sometime soon. And maybe you won’t run away screaming next time you see a radiation sticker. Genome editing is a vital tool for scientists to be able to understand diseases, improve crops, and unlock the secrets of how our cells work, as well as many other applications. Up until about 5 years ago, genetic engineering usually meant weeks of time and effort, was expensive and had limited versatility. Now, similar experiments can be conducted in less than a week very, very cheaply. So, what changed? The introduction of CRISPR/Cas9 technology revolutionised genome editing. CRISPR (clustered regularly interspaced short palindromic repeats) meant that genes could be knocked out (deleted), inserted, or mutated easily, with high accuracy, at almost any position in the genome. Based off of a bacterial immune defence system, the CRISPR/Cas9 complex consists of several components. The Cas9 protein forms a complex with guide RNA that matches the sequence of the DNA to be targeted. The complex scans the genome for the protospacer adjacent motif (PAM) sequence, which is a short series of DNA bases immediately after the targeted DNA sequence. When it finds a match, the guide RNA pairs with the target DNA sequence and the Cas9 protein cuts the DNA in half, forming a double stranded break. Double stranded breaks in DNA are bad news for the cell, so the cell attempts to repair the cut. However, this process is prone to errors, so often mutations are introduced which means the gene no longer functions. Alternatively, scientists can introduce new genes by adding DNA which is integrated at the site of the cut. In this way, scientists can add or delete genes to study how they work. CRISPR/Cas9 is very versatile and efficient, but it’s not perfect. While it can target almost any region of the genome, it still needs that PAM sequence. The most commonly used protein recognises NGG (N being any DNA base and G being guanine), which is relatively short but still has to be present in the target sequence. CRISPR/Cas9 is also limited by off-target effects. It may accidentally cut other regions of the genome that closely match the guide RNA sequence and introduce unwanted mutations.
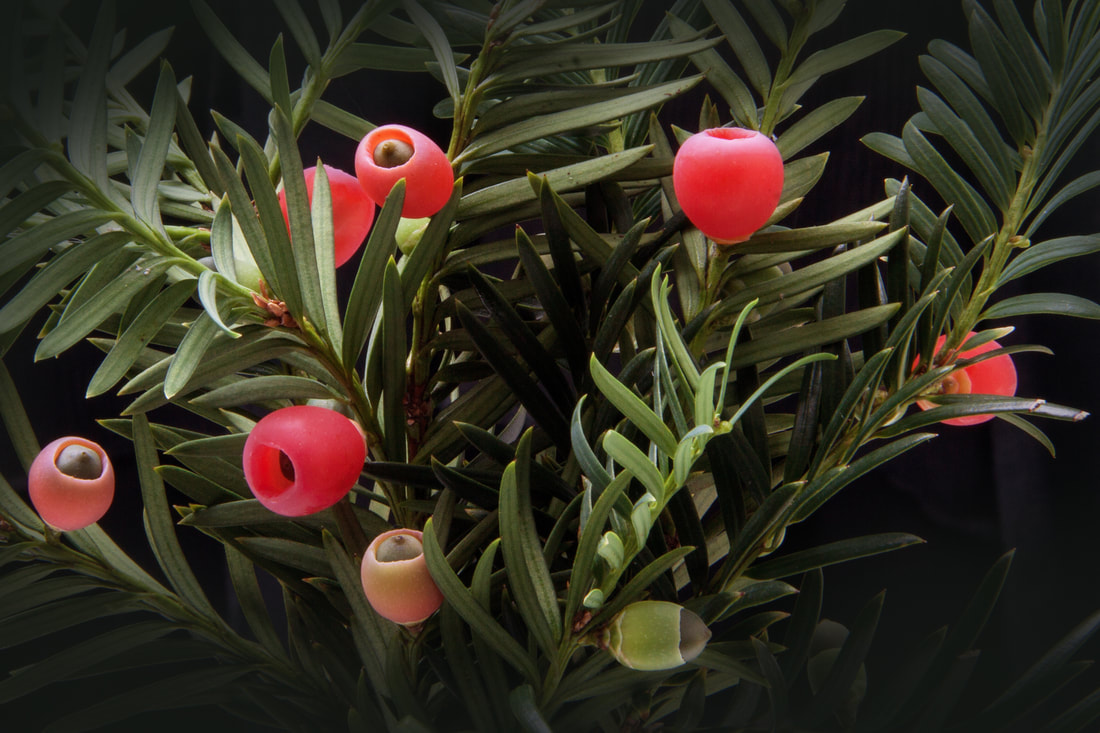
Recently, a group of researchers at Harvard set out to improve the Cas9 protein. They used a technique that forced the protein to rapidly evolve and accept a wider range of PAM sequences. They tested this new xCas9 variant with mammalian cells for compatibility and found that it will be suitable for a broad range of applications, especially in human cells. Surprisingly, xCas9 has greater specificity for the target DNA sequence, with fewer off-target effects observed than with the original Cas9. This illustrates that there is not necessarily a trade-off between specificity and versatility, and may help to improve future generations of Cas9 proteins. This improvement means CRISPR/Cas9 genome editing is even more versatile and will be able to target regions of the genome more specifically and more precisely, expanding the scope of genome editing and enhancing the potential of using CRISPR technology to understand and treat human disease. I got really excited about this research and just had to share it. I might be biased because it’s in my field of interest and uses some sexy maths, but I hope you enjoy it too. Many important compounds we use, like drugs and fragrances, are complex molecules. They’re often found in nature and produced by organisms like plants or marine life. Problem is, if we want a large amount of compound it can be hard to obtain. A good example is the cancer drug taxol (paclitaxel) that originally required killing up to four pacific yew trees to obtain one course of treatment. Thankfully, scientists have found a more efficient way to make the drug, but like many drugs and fragrances it still involves extraction from a natural source, which is expensive and wasteful. For some compounds, like the anti-malarial artemisinin, it also means the price of the product can fluctuate wildly depending on agricultural factors, demand and supply. Sometimes it is possible to make the product synthetically in a lab instead, but this depends on how complicated the product is and the availability of suitable starting materials. Clearly there is a need to develop better ways of making these important compounds.
Enter microbial biosynthesis. Microbial biosynthesis basically involves using a microbial organism, like yeast, bacteria, algae or fungi, to synthesise a useful product. This can be achieved from simple feedstocks (like corn syrup or plant by-products) through a fermentation process. The metabolic pathways of the organism are genetically manipulated to maximise production of the compound of interest, which could be anything from drugs and nutraceuticals, to fragrances and cosmetics, or even biofuels. Usually microbial biosynthesis is achieved using a single strain – that is one type of yeast or bacteria etc. This can be more efficient because it means intermediates don’t have to be transported across cell membranes but it has several downsides for complicated pathways. Firstly, the more complicated a pathway is, the more potential for metabolic ‘cross talk’ where the production of one intermediate unintentionally influences another metabolic pathway. Secondly, if an intermediate is toxic to the organism it will gradually cause mutations or death of high-producing populations leading to an overall lower yield of product. Thirdly, the burden of having to produce large amounts of compound in a single cell can slow down other metabolic processes meaning lower cell growth and ultimately less product. To get around this, an alternative is to use multiple cell types, such as combining two different types of bacteria, or a bacterial strain and a yeast strain, to share the load (Division of Labour). The idea is that one organism will produce an intermediate which is then used by the second organism to make the product. While this can reduce overall complexity by dividing tasks between the organisms, it can be limited by how well the intermediates are transported in and out of the cells. While division of labour has been used for individual cases, there’s no blanket rule or easy way to tell if it’s better to use one cell strain or multiple for any given project. Think about it like a team working on a project: sometimes it’s more efficient to just do all the work yourself rather than try and chase up people and pass documents around, but sometimes working as a team can lead to a better outcome. But how can you decide which way would be better? A team of researchers at Duke University have developed a model to understand the factors that influence productivity in a microbial system. They took into account things like enzyme production, rate of transport into and out of cells, rate of intermediate and product synthesis, and cell volume and growth to determine when it is more beneficial to use division of labour over a single strain. Effectively it boils down to two things: how fast the cells are growing and how well the intermediate is transported between the cells. From this, the researchers have come up with a general criterion to assess the efficiency of division of labour. Ideally, a scientist would be able to test these factors experimentally and use the model to decide which scenario is most appropriate for their project. Microbial biosynthesis is still relatively in its infancy in terms of industrial applications, but as technology improves, with the help of modelling systems like this one, more efficient syntheses can be achieved for the cheaper and more environmentally friendly production of important molecules. |
Author
Emi Schutz Archives
March 2018
Categories |











 RSS Feed
RSS Feed
