|
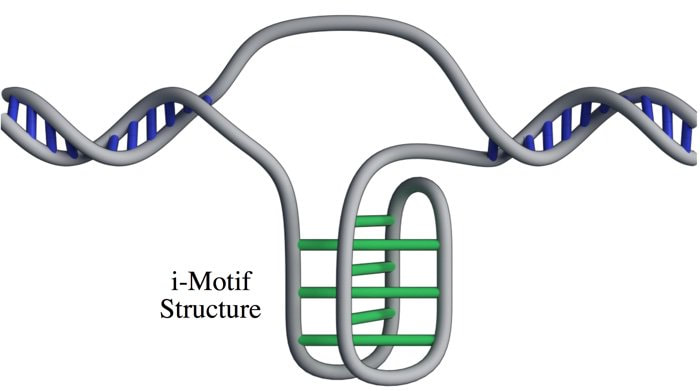
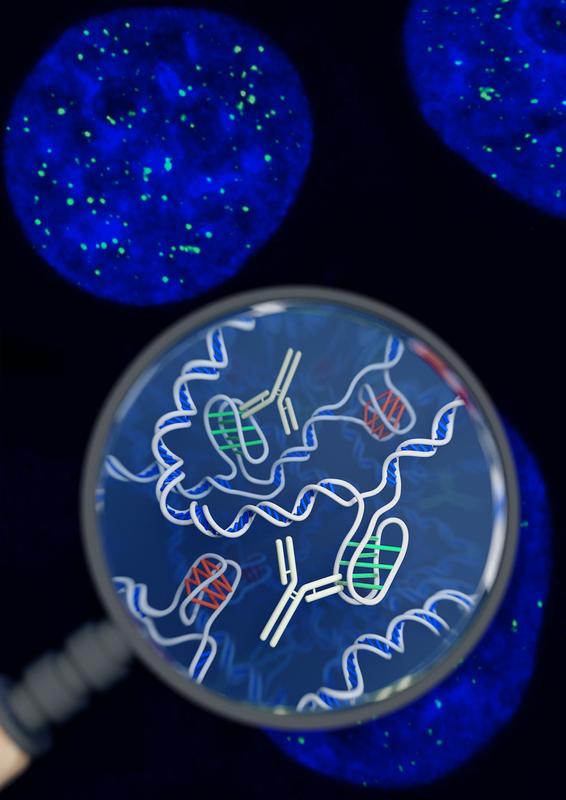
Most of us are pretty familiar with the double-helix structure of DNA; the one that looks like a spiral staircase with two strands of DNA pairing to each other. But did you know DNA can come in lots of other shapes too? DNA can twist and turn in different directions, pair up differently, and even form a triplex structure. Studying the structure of DNA is important because it gives insights into how DNA is used and read within the cell. Our DNA is really long, so it has to be packed in pretty tightly to fit inside the nuclei of our cells. In order to read the DNA to express genes it has to be unpacked and unwound, and there are lots of different proteins and big molecules (like RNA) that come into play in this process. Understanding the different structures of DNA and how they regulate gene expression can tell us information about what is happening in the cell and how that might impact health and disease. One structure – the i-motif – looks a bit like a knot of DNA. Normally the DNA helix structure is formed by the pairing of base letters A, T, C and G on two different strands of DNA, but the i-motif ‘knot’ structure is formed by the C-bases on the same strand of DNA binding to each other. (Zeraati et al., Nat Chem, 2018) This had only ever been observed in vitro, that is, outside of the human body, and usually only under harsh pH conditions. While there was some indirect evidence for its formation in vivo, it wasn’t known whether or not it actually existed in living organisms. This week, Australian scientists announced they had found a way to detect this i-motif inside living human cells for the very first time. Their findings were published in the journal Nature Chemistry. The researchers developed an antibody that binds specifically to the i-motif structure. They did this through three rounds of identifying and refining a mutant antibody that could bind to a known i-motif structure found at the end of chromosomes. They found the antibody bound strongly and specifically – that is, it didn’t interact with other proteins, RNA, or other DNA structures. They tested the antibody on six other known i-motif structures and found it was able to bind to all of them, indicating the antibody can bind to i-motifs regardless of the DNA sequence. That meant it could be used for detecting i-motifs inside cells. The researchers stained human cells using a technique that makes fluorescent spots in places where the antibody binds. They found multiple fluorescent green dots, showing for the first time the presence of i-motifs in living cells. They observed the i-motifs appearing and disappearing over time as the cells grew and divided, and found they were linked to regulatory regions of DNA. This research not only gives direct evidence for i-motifs in cells but also provides preliminary evidence that they could be involved in human diseases that arise from dysregulation of gene expression, such as cancer. Mutations in i-motif regions could change the stability or shape of the i-motif and affect how it interacts with DNA. This research, and the new antibody tool, opens new doors for future studies to explore the biological roles of i-motifs and potentially develop new therapeutics to treat disease. Artist's impression of the i-motif DNA structure inside cells, along with the antibody-based tool used to detect it. (Chris Hammang).
0 Comments
Leave a Reply. |
Author
Emi Schutz Archives
March 2018
Categories |


 RSS Feed
RSS Feed
