|
How many times have you watched a murder mystery show where the detective strolls into the murder scene and inspects the victim’s body for just a few moments before confidently declaring the time of death? This may not come as a surprise, but in real life it’s not that simple. In the first few hours, time of death is usually estimated by taking into account physiological factors, like body temperature, rigor mortis and skin discolouration. However, these methods are still largely unreliable and inaccurate, as they can be influenced by a number of factors including the ambient conditions, cause of death, body structure, body location, and drug consumption. Researchers have been searching for ways to make estimating time of death more accurate, and one of these new methods involves looking not just at the body as a whole, but analysing what is inside cells at the molecular level.
Our genetic information is stored in cells as DNA. In order for that information to actually do something, in needs to be read. When DNA is read it is transcribed into RNA, which is then translated into proteins that undertake some action in the cell. When a tissue dies, the cells in that tissue undergo changes in gene expression. A team of researchers in Portugal and Spain has looked at how gene expression changes after death by analysing RNA levels in pre-mortem and post-mortem samples. They found that each tissue exhibits specific changes in gene expression that is not consistent with random RNA degradation but actually a result of active and ongoing gene regulation, even after the organism dies. These changes are speculated to be caused by low oxygen as the blood supply to the tissue is lost. Each tissue type has its own unique pattern of changes in gene expression over time since death. From this, the researchers built a model that can be used to estimate time of death by looking at RNA levels in tissues that are likely to be found in a forensics scenario, including fat, lung, skin and thyroid tissue. They also found no impact of cause of death on their estimates. This technique could not only improve time-of-death estimates in forensic pathology, but also assist researchers working with post-mortem tissue samples and have implications for organ preservation and transplantations. Also, if you’re curious as to how forensic scientists determine time of death after days, or even weeks and months, check out this article on forensic entomology.
0 Comments
The world population is predicted to reach 9.8 billion by 2050. In order to feed this number of people, we need to dramatically increase cereal yields, above what is already being achieved by plant breeders. Wheat is one of the oldest and most important cereal crops. Without it we wouldn’t have tasty food like bread, pasta, cake, and pizza. (I get hungry just thinking about wheat. So delicious). Some of the factors that can be selected for to improve wheat yield include nitrogen uptake, leaf mass and photosynthetic efficiency. However, in order to select the best plants for breeding, all of these factors need to be measured. Traditionally these measurements are time consuming, destructive, and require special facilities, which is not practical on the large scale required for plant breeding. Ideally, researchers want to be able to tease out genetic markers they can look for that will tell them if a plant, or its offspring, will be high-yielding. However, considering the wheat genome has 6 times more base pairs than humans, this is no easy task!
Recently a group of researchers at the ARC Centre for Excellence for Translational Photosynthesis sought to speed up the wheat-analysis process. They wanted to see if hyperspectral reflectance– that is reflecting light in colours beyond the visible spectrum, like UV and infrared – could be linked to photosynthetic- and biomass-related traits. To test this they grew a lot of wheat plants in a greenhouse and took measurements of traits like carbon dioxide conversion, leaf mass, nitrogen content and chlorophyll content, as well as the leaf reflectance over a full range of wavelengths (350 – 2500 nm, or UV-Vis-Infrared). They then put these measurements together to construct a statistical model of how well hyperspectral reflectance can be linked to the measured traits. Their model is able to provide robust estimates for 6 traits that are linked to wheat yield, biomass and photosynthetic ability. The importance of this technique is that it is very fast compared to traditional methods. It used to take researchers 20 minutes per leaf to obtain this kind of information, but using hyperspectral reflectance approximately 100 plants can be analysed within an hour. Higher-throughput will make it easier to screen large populations for breeding and to identify genetic markers that will increase wheat yield. The researchers hope to expand their model by adding more leaf properties to measure, and the technique is starting to be used for other crops including corn, rice and sorghum. Down the track, using hyperspectral reflectance will hopefully lead to higher grain and cereal yields to meet the demands of a growing population. Whether you say ‘lay-go’, ‘leg-oh’, or ‘lee-go’, we can all agree that LEGO is pretty awesome. Personally, my childhood was filled with LEGO houses and castles, tower building competitions, and shredded fingernails from trying to prise apart small plates (did you know they have a tool for that now?). One of the best things about LEGO is that you can create almost anything, from giant sculptures to a working mechanical keyboard. But who knew LEGO could be so versatile that it could be used to build a tiny, modular, microfluidics lab? Microfluidics is basically the manipulation of fluids on a really small scale. Fluids are moved, mixed, separated or otherwise processed through tiny capillaries on the sub-millimetre scale. This can be used for things like enzymatic analysis, DNA sequencing, continuous testing for pathogens or toxins, and many other biological, optical and chemical applications. Microfluidic devices usually take the form of a ‘lab on a chip’ – a tiny, flat surface etched with channels and ports to manipulate the flow, mixing, and separating of the fluid. However, the problem with this is that so far there’s no universally accepted way to construct this device, meaning each one needs to be custom synthesised for its purpose, and it’s hard to mix and match devices. On top of that, they’re not exactly easy to make either as 3D printing is not yet precise enough for this purpose so construction usually requires expensive facilities, materials and labour. An alternative method is to use a modular system, that is, each module performs a single function. And scientists at MIT have found a novel platform: LEGO bricks. In their recent paper, published in Lab on a Chip (cool journal name btw), Crystal Owens and John Hart explored the use of LEGO bricks to build a microfluidic platform. They went out to the shop and bought an off-the-shelf LEGO set, and used a micromilling technique to precisely etch tiny channels into the bricks. To seal the faces, the bricks were covered in a thin plastic film, and brick modules were joined via an O-ring. Because LEGO bricks are so consistent in size and fit together easily, the O-rings formed reliable and reversible seals that allow the microfluidic device to be literally ‘clicked’ together. While this system is cheap to build and extremely reliable, it isn’t perfect yet. Firstly, the micromilled channels are relatively large and not suitable for many applications. They’re also made of plastic so they wouldn’t be suitable to use with some organic solvents. Future work will look at different types of materials and different ways of constructing the bricks so they can be used for a wider range of applications. But for now, a LEGO ‘lab on a brick’ could be the future of prototyping these tiny microfluidic devices. Here’s a fun fact: approximately 1 in 10 people in the world live within 100 km of an active volcano. Aside from causing mass evacuations, destroying huge areas of land, and ruining your travel plans, volcanic eruptions can have devastating health effects. Some research estimates about 540 people are killed by volcanic activity each year. At the moment, there’s no accurate way to predict when a volcano will erupt, but that could change thanks to the work of some Queensland scientists. To understand how this research works, first let’s talk volcanoes 101. Imagine the world like a Ferrero Rocher: a hard crust on top of a viscous mantle with a solid core in the middle (ok, there’s also a liquid inner core but that doesn’t fit with my analogy). Credit: A. Kniesel, Wikimedia CC BY-SA 3.0 The crust is relatively thin, like the skin on an apple, and it is broken up into tectonic plates that sit on the mantle. Volcanoes tend to occur where these tectonic plates meet and either run into each other or move away from each other. Weaknesses in the crust allow molten magma from the mantle to be pushed up, forming a volcano. Within the volcano, magma collects in magma pools, until the build-up of pressure from molten rock and gas forces the magma upwards through weaknesses in the surrounding rock. Eruptions occur when the pressure is so great the magma is pushed to the surface and the volcano explodes, releasing lava, ash and gases. A sudden influx of new magma into existing magma pools can be a trigger for a volcanic eruption. However, since each volcano is different and has complicated pathways from magma pools to the surface, it can be hard to tell how long it will take from a magma influx to an eruption for each individual volcano.
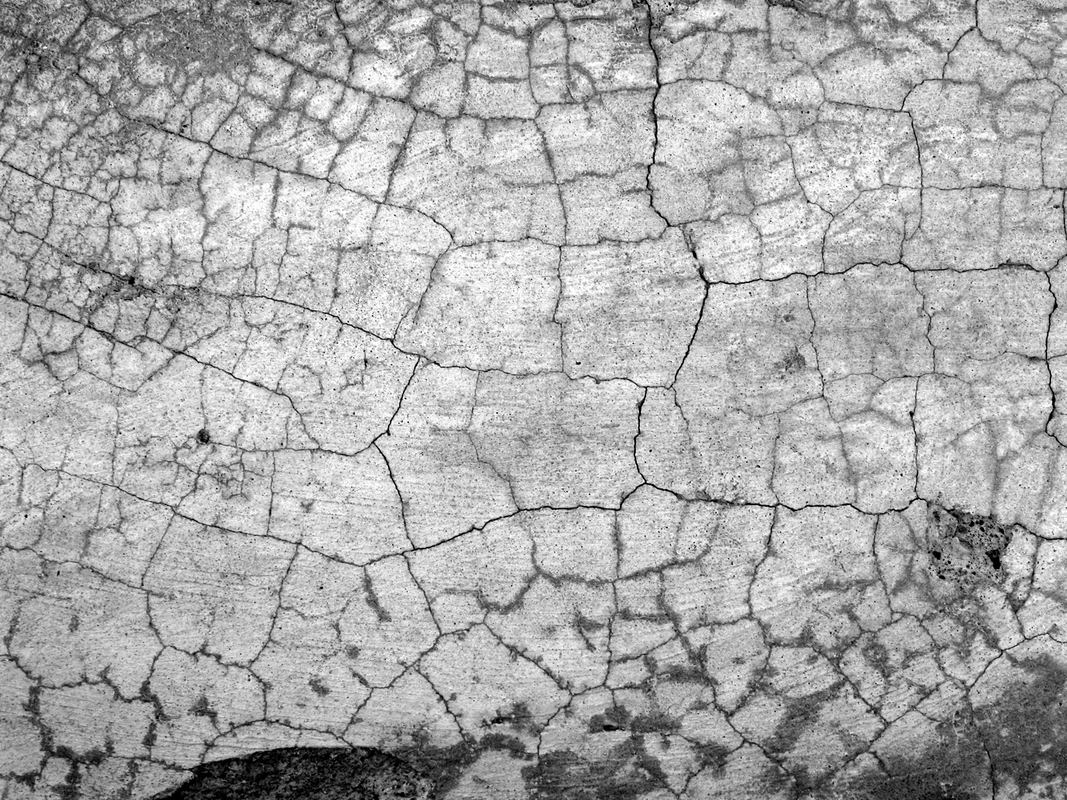
By looking at the chemical composition of tiny crystals and matching this with geophysical and eruption data from the past 40 years, a team of researchers at the University of Queensland has built a model to predict the eruptions of Mt Etna (Sicily, Italy) with a success rate of up to 90%. The crystals grow as magma starts to move up towards the surface of the Earth. The composition of the crystals change depending on their surrounding environment, and grow like tree-rings. By mapping the trace elements within the crystals, the researchers found chromium-rich zones, which suggest crystal growth shortly before eruption, that were surrounded by a chromium-poor ring, indicating surface crystallisation after eruption. By working on the premise that chromium-rich zones indicate the influx of new magma the researchers were able to suggest pathways for the flow of magma through Mt Etna, based on regular eruption events over the past decades. Based on these pathways and seismic data it is possible to predict that Mt Etna will erupt within 2 weeks of a magma influx. The researchers hope to apply this technique to other active volcanoes to better predict when eruptions will occur, allowing early evacuation, saving lives, and not ruining your next trip to Bali. Cracks in concrete are unsightly and expensive. They start off small but they can grow pretty quickly, and if water gets in the steel reinforcing underneath can be damaged and lead to structural failure. You can replace the concrete, but then it’s only a matter of time before cracks form again. So wouldn’t it be great if the concrete could just…heal itself? This idea is not actually as far-fetched as it might seem. Self-healing concrete is not new, and previous research has tried to accomplish this via three main processes. The first is autogenous healing. When water flows into a crack, cement particles in the crack are hydrated and calcification occurs due to dissolved carbon dioxide in the water. However, this only works for very small cracks and requires water. The second is the incorporation of a polymer matrix into the cement. When the polymer is exposed to humidity it releases a foam-like substance to fill the crack. Unfortunately, since this material behaves very differently to the concrete it can sometimes only make the cracks worse. The third method is bio-based: using calcium carbonate-producing bacteria to patch up the concrete. Bacteria are pretty good at this, but there are a few drawbacks, like increased nitrogen being released into the environment and needing a highly concentrated calcium source.
But now there’s a new kid on the block: fungi. A team of researchers at Binghamton University, State University of New York conducted an experiment analysing the interactions of various species of fungi with concrete. They found a fungus called Trichoderma reesei that is able to produce calcium carbonate, and can survive the drastic pH change that occurs as concrete dissolves. The idea is that this fungus will be mixed into the concrete. It will remain in a dormant spore state until a crack appears, exposing the spores to water and oxygen. When enough water and oxygen seep in, the fungus will wake up and begin growing and multiplying. As it does this it will precipitate calcium carbonate to fill up the crack. Once the crack is filled and no more water and oxygen can enter, the fungus will return to a spore state. The research is still in its early stages and the main challenge now is to make sure the fungus can survive within the concrete and throughout the concrete-making process, but bio-based concrete-healing is definitely a new technology to look out for. Self-healing concrete – coming soon to a neighbourhood near you? Imagine a world where a minor injury or infection could be a death sentence. It sounds like the dark ages, the pre-antibiotic era before WWII, but it could once again become reality. The world is fast running out of effective antibiotics. Bacteria have evolved resistance to all known antibiotics, and the prevalence of antibiotic-resistant infections is on the rise. The problem is so bad that the World Health Organisation predicts that in 2050 10 million people will die from antibiotic-resistant infections. Source. Image Credit: Katie McKissick Used with permission. Part of the problem is that it’s really hard to find new antibiotics. Many are based on natural compounds that other bacteria produce as a defence mechanisms, and here all the low-hanging fruit has been picked. Purely synthetic (man-made) compounds are another avenue, but often compounds that seem initially promising are later found to be ineffective because they can’t pass through cell membranes to actually kill the infection. Oh, and there’s never enough funding. Like with all drugs, antibiotic discovery has a high failure rate. The best way to get around this is to try lots and lots of compounds. I’m talking hundreds of thousands. No, not hundreds and thousands
To overcome these limitations, a team of researchers at The University of Texas at Austin has come up with a new technique to screen potential antibiotics: Surfaced Localised Antimicrobial Display aka SLAY (because everything in science needs a cool acronym, even if slightly contrived). Here’s my three favourite things about the technique:
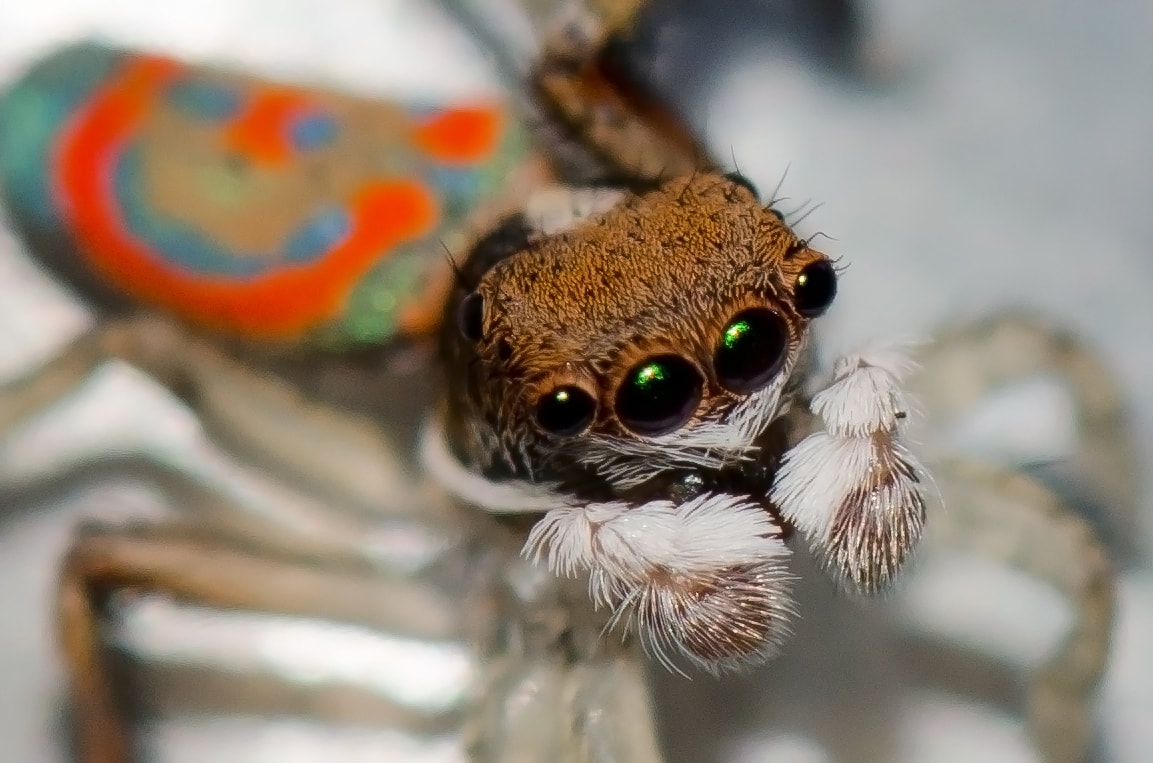
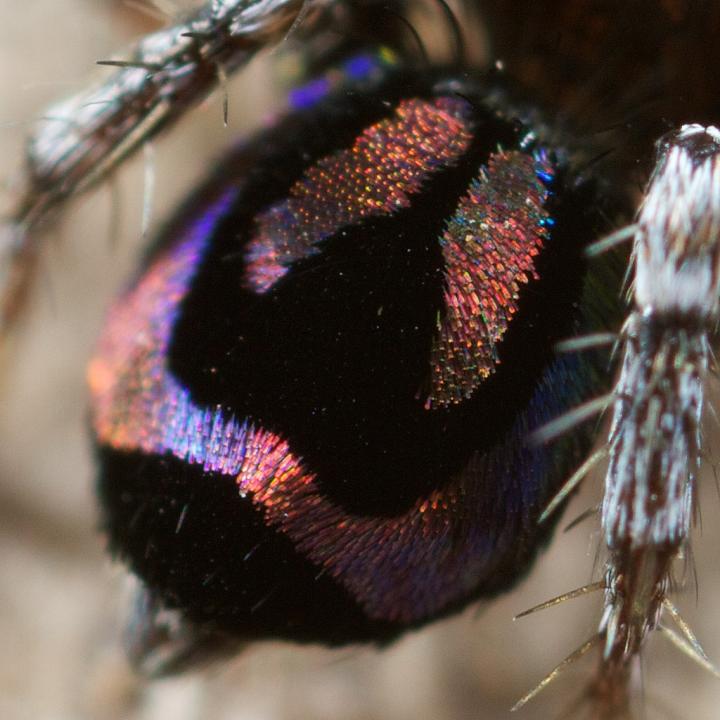
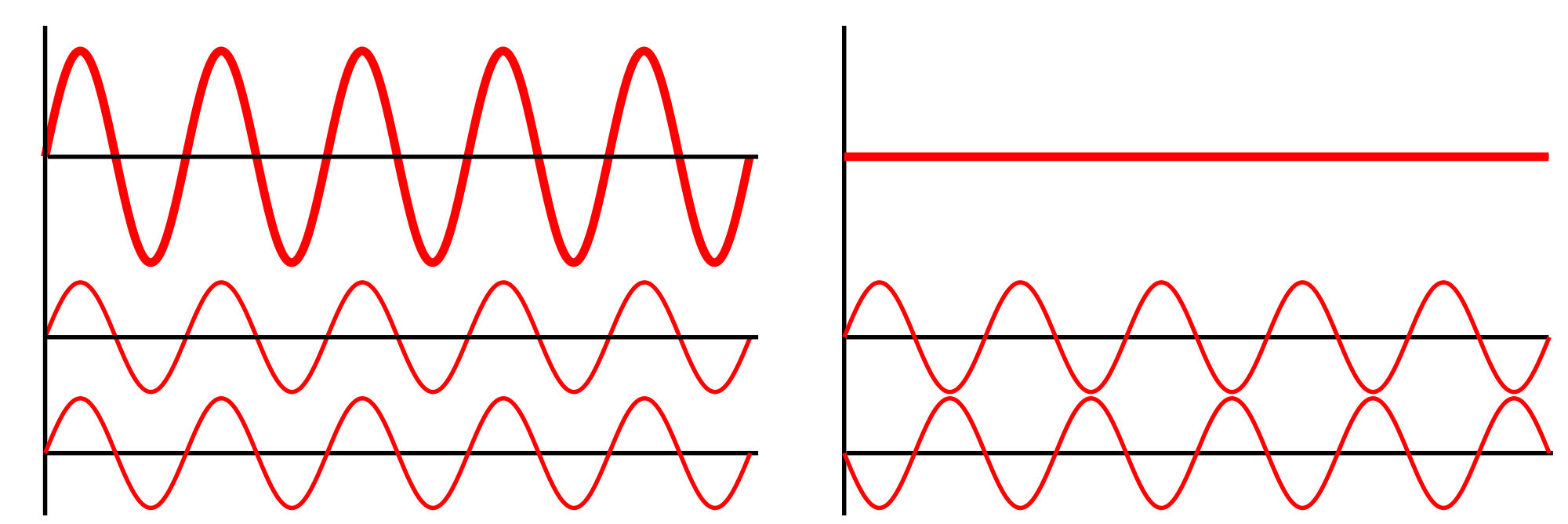
Of the 800,000 random sequences, the team found 7,968 potential new antimicrobial peptides. They tested 22 of these against four different strains of bacteria and approximately 80% were active against the bugs. They’re now working on developing these active compounds further to make them even more effective. With any new compound it is only a matter of time before the bacteria evolve and catch up. Antibiotic resistance is inevitable. Like in the And the scientists working on it? I bet they SLAY all day. I know not everyone is a huge fan of spiders, but I think this guy is pretty cute: Photo credit Jean and Fred, Flickr. [CC-BY 2.0] He’s a Peacock Spider (Maratus spp.) and his way of impressing the ladies is to do a little dance, showing off his brightly coloured backside: As he moves, notice how the colours change all the way from purple to blue through to yellow and red. In fact, this little spider can show off all colours of the rainbow, depending on which way you look at him. Iridescence in nature is not uncommon – think of butterflies, peacocks and pigeons – but usually it’s over a very limited range of colours, like a blue that shifts from bluey-green to purple. It turns out this little spider is incredibly unique because he’s able to display every colour in the visible spectrum. So, how do these spiders make their rainbows? In a recent paper published in the journal Nature, researchers set out to answer just that. They used a range of imaging and probing techniques including electron microscopy and optical modelling to figure out how the spider scales display such beautiful colours. To understand iridescence we need to think about light as waves that can interact and interfere with one another. If two waves meet so their peaks line up, the peaks will add together and amplify the wave, making it twice as big. This is called constructive interference. On the other hand, if a peak meets a trough it will cancel out to nothing, which is called destructive interference. Iridescence can be caused by reflecting light multiple times over several semi-transparent layers, or by using a diffraction grating which splits incoming light into several beams travelling in different directions. In both of these cases it is constructive interference that creates iridescence. The researchers found the spiders have a microscopic 3D surface that is curved a bit like an aerofoil. Over this surface they have a nanoscale diffraction grating. The interaction between the grating and the curved surface means that light hitting the scale is separated out into its individual colours resulting in a beautiful rainbow.
The researchers then used nanoscale 3D printing to try and mimic the spider’s intricate structures in order to confirm their hypothesis. Our current technologies are not capable of resolving and separating white light into individual colours at short distances, but inspiration from these spiders’ scales could help improve optical technologies. In particular it could reduce the size of spectrometers used in space missions, or even lead to wearable chemical detectors. Who knows, at some point in the future we could all be using spider-inspired technology! “New year, new me” – every teenage girl on Instagram* The New Year seems like an ideal time to make resolutions because it feels like a fresh start. We are drawn to important dates, like the start of a new year, new month, or even a new week, to begin our goals. But how many of us fall into the trap of making the same New Year’s resolutions over and over again? And how many of us scoff when other people tell us of their grand goals and ambitions, certain that their plans will have fallen by the wayside in mid-January? I never keep my resolutions so this year I’m going to eat more chocolate, exercise less and spend more money. It’s one thing to make resolutions, but another thing entirely to keep them. Since my New Year’s resolution is to blog once a week, I’m starting with five science-backed ways to keep your New Year’s resolutions.
Perhaps I should only let myself eat doughnuts while writing my blog? What’s your new year’s resolution (and how are you planning to achieve it?). I’d love to hear from you – please leave a comment below. If you have found any of these tips interesting or useful, please share with a friend. This is only a little blog in a big, big internet, but I’d love to get my writing out there for everyone to enjoy. On that note, goodbye 2017, and bring on 2018! *If you are a teenage girl on Instagram, I sincerely apologise. I’m sure you’re an exception. Acknowledgements: The idea for this article and some of the supporting research was inspired by the Hidden Brain podcast. I’m new to the world of social science research and the podcast gave me a great starting point to find some of the above papers. If you’ve ever wondered whether the way you park says something about you, or how to teach monkeys to use money, check out their podcast here. Hello and welcome to my brand new blog! I started SciChat as my 2018 New Year's Resolution to write more often. Each week I'll find an interesting recent research paper and write about why I think it's so fascinating. I'll occasionally post bonus content as well if I have time. I don't claim to be an expert in every (or any) area of science but I have a pretty solid grounding in physics, chemistry and biology. I can't guarantee I'll get everything right though so I always welcome feedback on my posts and like any good scientist I'm willing to reconsider my viewpoint if new evidence arises.
I know blogs can be a little bit 2008 and it might just be my Gran reading this (hi Gran!) but I do hope you keep coming back here, or subscribe to RSS updates or follow me on Twitter to keep up with my latest posts. I can't promise this is the next IFLScience but I hope you find my posts entertaining and learn something new each time. Finally, thanks to all the generous artists who posted their images on Pexels and Unsplash. This is where I sourced most of the header and background images for my site and I think they are absolutely stunning (and best of all, free). I promise to have my next post up by New Year's Day, so stay tuned! |
Author
Emi Schutz Archives
March 2018
Categories |












 RSS Feed
RSS Feed
